Welcome to GLG Pharma
Our Technologies for Cancer are Unique.
Get started with us to learn how.
Great Drug Discovery is Reliant on Reproducible, Relevant, and Actionable Results from the Lab.
Isolating the Wrong Cancer Cell Clonal Populations Leads to Clinical Drug Failure
Tumor and Patient Heterogeneity and not being able to isolate and identify all the cancer players; Drivers, Passengers, Sleepers, Enablers, and Ambassadors means your drugs and drug combinations will not survive the clinical trials process, no matter how much you and the team know about the mechanisms of action, binding, distribution, efficacy, and safety. Heterogeneity will win.
- Use the right tools in the lab for the media to replicate the patients tumor signatures, all of them.
- Understand the sub-clonal and clonal populations and what structural architecture they like
- Use Single Cell DNA and RNA information in Drug Discovery to parse the various players in the cancer Tumor microenvironment
What We Do Sets Us Apart

Designing Media for the Cancer
Leverage the 20+ years of cancer cell culturing knowledge imbedded in our cancer cell culturing media. Saves you time and will rescue your research and your business. Promise.

Available Media for Cancer Cells
Leverage the awesome cancer cell culturing media that saves you time and will rescue your discovery research. Promise.

Partnering
Leverage the combination of cancer cell technologies and drug discovery for your research projects. Partnering doesn't have to be difficult and it can save money. Promise.
An Example of Our Media for Cancer Cell Culturing
Epithelial Example: Ovarian Tumor Cells
In xenographs, morphology is also better preserved
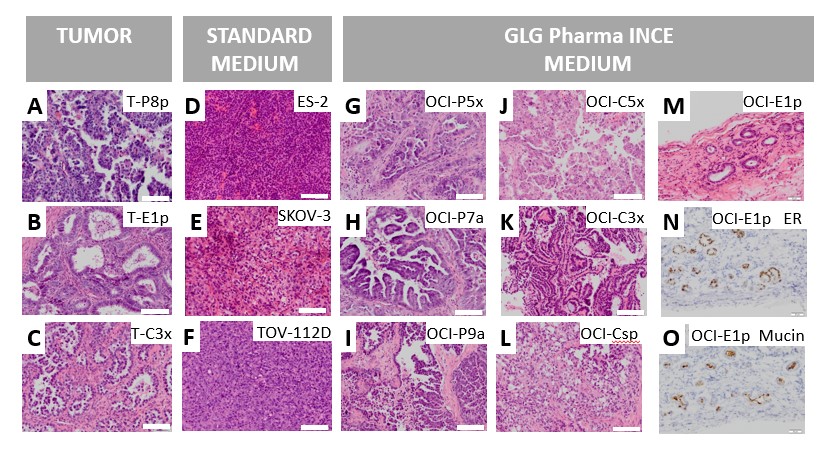
Why is GLG Pharma Different?
Our Approach Works
Let our tested and proven approach to culturing cancer cells help you and your team identify the best technologies, diagnostics, and drug candidates whether small molecule or large biomolecules, using the right media for the right cancer cells is the key.
Match to Patient's RNA/DNA tumor profiles in lab cancer cultures
Increase increase in the ability to culture ovarian cancer cells in the lab using the GLG Pharma INCE medias

